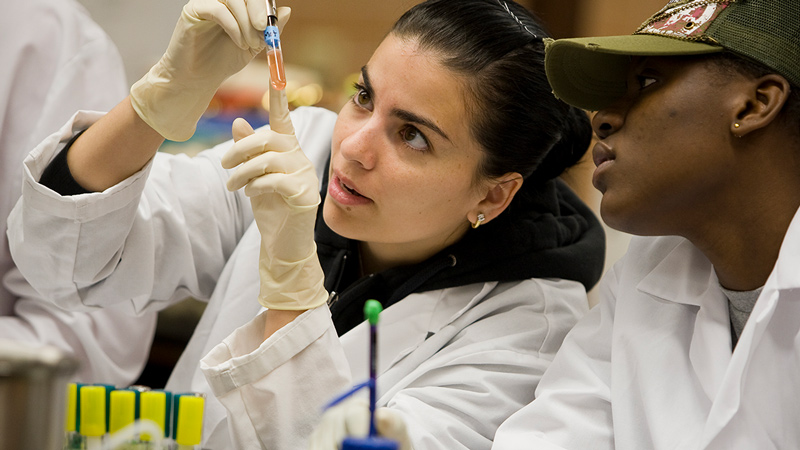
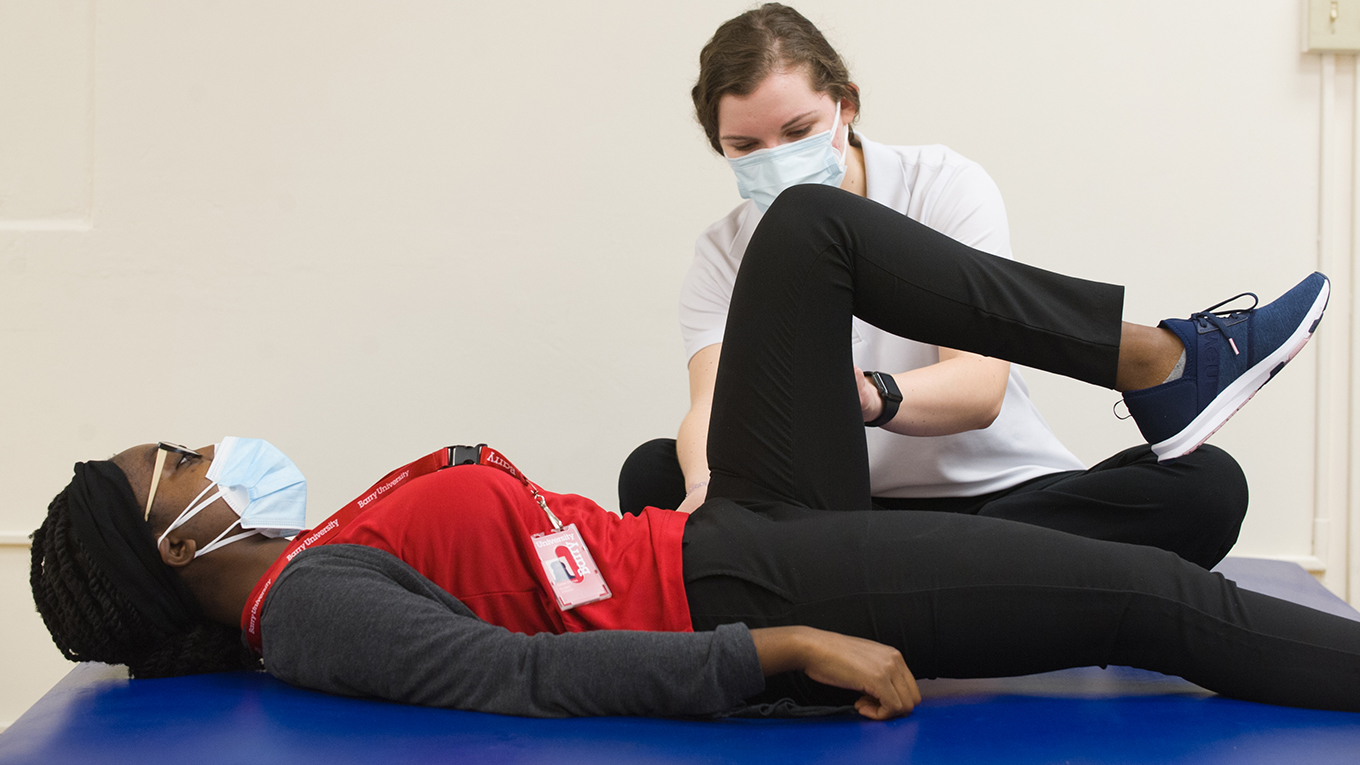
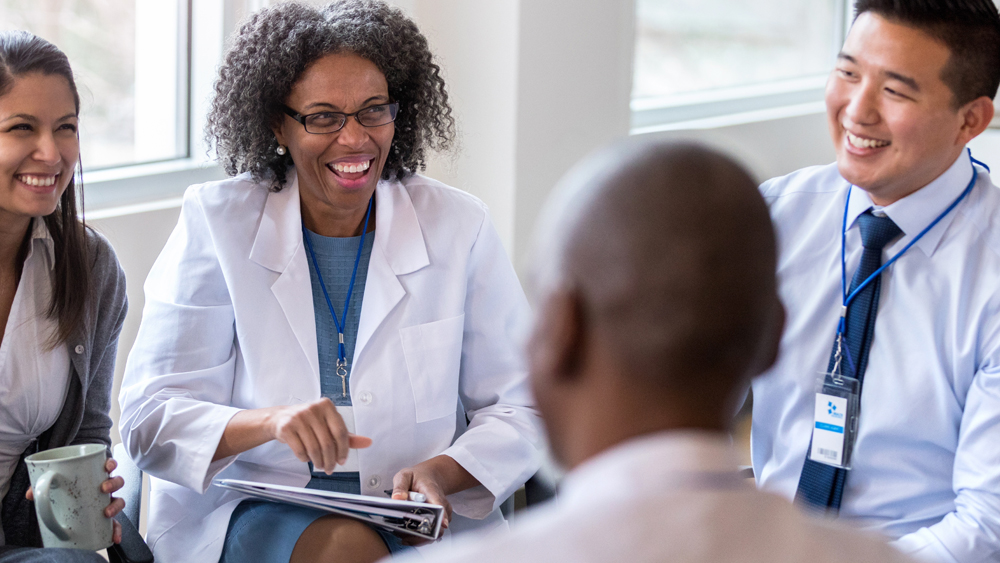
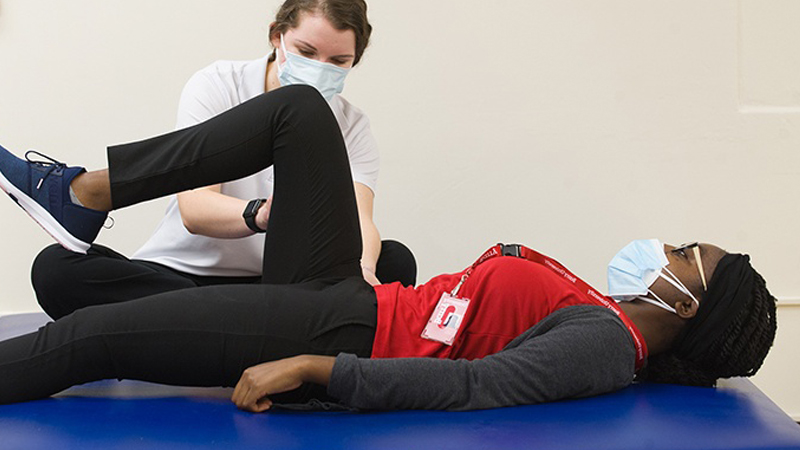
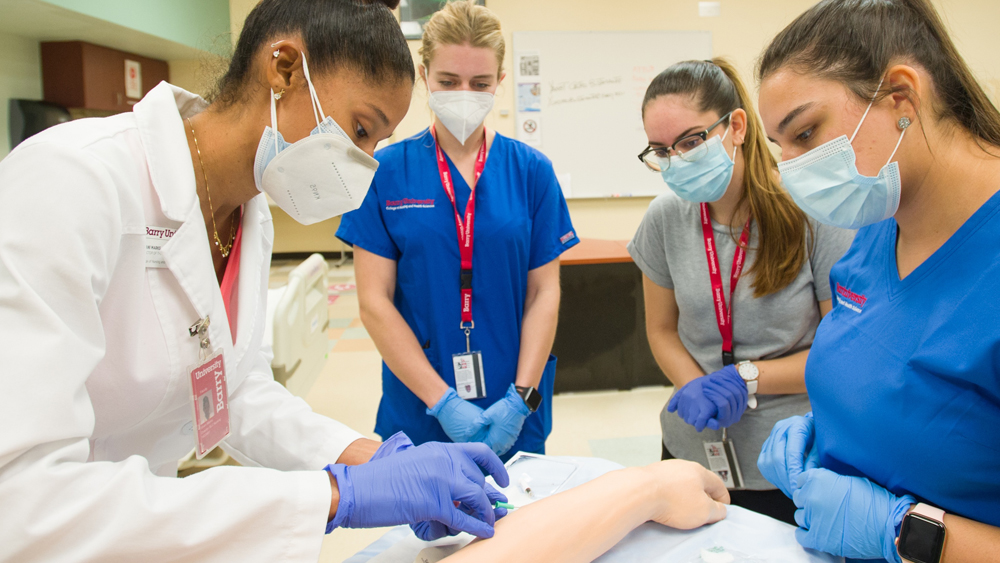
Your journey to success in today’s evolving healthcare environment starts here.
Our Programs
Browse all our programsChoose your College
Discover your path to success.
- Decades of leadership and respect in preparing healthcare professionals.
- Hands-on, highly regarded faculty with strong clinical backgrounds.
- High employment rates and proven outcomes from patient care and diagnostics, to community service, to high-ranking corporate positions.
- A nurturing faculty that represents and respects diversity and inclusivity.
- An integrated and collaborative learning environment across all healthcare professions.
- Compassion, ethics and inclusivity are intrinsic throughout our courses.
- Seamless programs that will help you move from entry level to advanced degrees.
- Ample opportunities for research, and hands-on clinical practice at renowned South Florida hospitals.
You can expect excellence from our faculty. And we will expect excellence in your health care practice.

Top Stories
Locations Miami is not the only location!
Main Campus
Address
11300 NE 2nd Ave Miami Shores, FL 33161
Phone
305-899-3000St. Petersburg Pinellas
Address
9200 113th Street N Seminole, FL 33772
Phone
305-899-4035St. Petersburg Physician Assistant Program
Address
7200 66th Street North Pinellas Park, FL 33781
Shands at University of Florida Clinical Site
Address
2000 SW Archer Road, Suite #4164 Gainesville, FL 32608
Phone
800-556-3932Broward General Medical Center Clinical Site
Address
1600 South Andrews Avenue Fort Lauderdale, FL 33316
Phone
954-355-4400Broward Health Imperial Point Clinical Site
Address
6401 North Federal Highway Fort Lauderdale, FL 33308
Phone
954-776-8500Broward Health Coral Springs Clinical Site
Address
3000 Coral Hills Drive Pompano Beach, FL 33065
Phone
954-344-3000Broward Health North Clinical Site
Address
201 E Sample Rd Pompano Beach, FL 33064
Phone
954-941-8300Tampa General Hospital Clinical Site
Address
1 Tampa General Circle Tampa, FL 33606
Phone
813-844-7000Moffitt Cancer Center & Research Institute Clinical Site
Address
12902 Magnolia Drive, SRB4 - BMT Tampa, FL 33612
Phone
888-820-0230Advent Health Ocala Clinical Site
Address
1500 Southwest 1st Avenue Ocala, FL 34471
Phone
352-351-7200Aventura Hospital and Medical Center Clinical Site
Address
20900 Biscayne Blvd Aventura, FL 33180
Mount Sinai Medical Center Clinical Site
Address
4300 Alton Rd Miami Beach, FL 33140
Phone
305-674-2273Nemours Children's Hospital Clinical Site
Address
13535 Nemours Pkwy Orlando, FL 32827
Phone
407-567-4000Osceola Regional Medical Center Clinical Site
Address
700 W Oak St Kissimmee, FL 34741
Phone
407-846-2266Good Samaritan Medical Center Clinical Site
Address
1309 N Flagler Drive West Palm Beach, FL 33401
Phone
561-655-5511Jupiter Medical Center Clinical Site
Address
1210 South Old Dixie Highway Jupiter, FL 33458
Phone
561-747-2234Palms West Medical Center Clinical Site
Address
13001 Southern Blvd Loxahatchee Groves, FL 33470
Phone
561-798-33001700 South Tamiami Trail Sarasota, FL Clinical Site
Address
1700 South Tamiami Trail Sarasota, FL 34239
Phone
800-764-8255Doctors Hospital of Sarasota Clinical Site
Address
5731 Bee Ridge Rd Sarasota, FL 34233
Phone
941-342-1100Academics Healthcare Leaders are Made Here
Barry University’s guiding principles to study, reflect and serve the community are evident throughout our wide variety of programs. Our goal is to prepare inspired and compassionate healthcare leaders in the delivery of high quality, accessible, culturally discerning care within an ever-evolving, highly technological, global environment.
The Barry Touch makes us different – while many university programs can prepare you intellectually, our faculty trains your hands to touch with compassion and expertise, trains your heart for the ongoing care for others, and trains your mind to react with agility. As patients and hospitals can attest, there are healthcare providers, and then there are Barry providers. The very best healthcare practitioners and providers are made here.
Contact Us
We’re always happy to chat with you about your career goals and the academic programs to get you there. Our team of admissions experts offers many communication options from chat or text to online meetings and email options.
Get all your questions answered at a convenient virtual info session including application assistance.
College of Health and Wellness Our Faculty are Passionate Healthcare Experts
At the College of Health and Wellness at Barry University, you’ll be inspired, guided and pushed to excel by a vibrant and dynamic faculty who are at the forefront of society’s evolving healthcare needs. We seek out professors who are not only great teachers, but who love what they do. As working practitioners, our faculty members maintain positions at hospitals and clinics, or have independent practices – keeping them fully up to date on the latest technology, patient care practices and legislation.
Beyond each teacher's area of expertise, our faculty is committed to promoting equality, social justice, altruism, diversity, and humility. Here, you will experience The Barry Touch. Our faculty will train your hands to touch with compassion and expertise, will guide your heart for the ongoing care for others, and will train your mind to react with confidence and agility. In the skillful hands of our respected faculty, will you become an extraordinary healthcare leader.
College of Health and Wellness Accreditation
Barry University is accredited by the Southern Association of Colleges and Schools Commission on Colleges (SACSCOC) to award baccalaureate, masters, specialist, and doctorate degrees.
Questions about the accreditation of Barry University may be directed in writing to the Southern Association of Colleges and Schools Commission on Colleges at 1866 Southern Lane, Decatur, GA 30033-4097, by calling (404) 679-4500, or by using information available on SACSCOC’s website (www.sacscoc.org).

Support our Health & Wellness Students
There are many ways to support our students in the College of Health and Wellness. Whether it is to help students succeed through academic and clinical experiences, enhance the simulation environment, or invest in latest technologies in physical and psychological patient care.
Your gifts are an investment in the future of healthcare. Every philanthropic investment makes a lasting impact and supports our healthcare and wellness programs.
Interprofessional Collabration
At Barry University we take pride in offering our students the opportunity an interprofessional collaborative approach to develop healthcare students as future interprofessional team
members. Training future healthcare providers to work in teams will help to facilitate improved healthcare outcomes for patients.